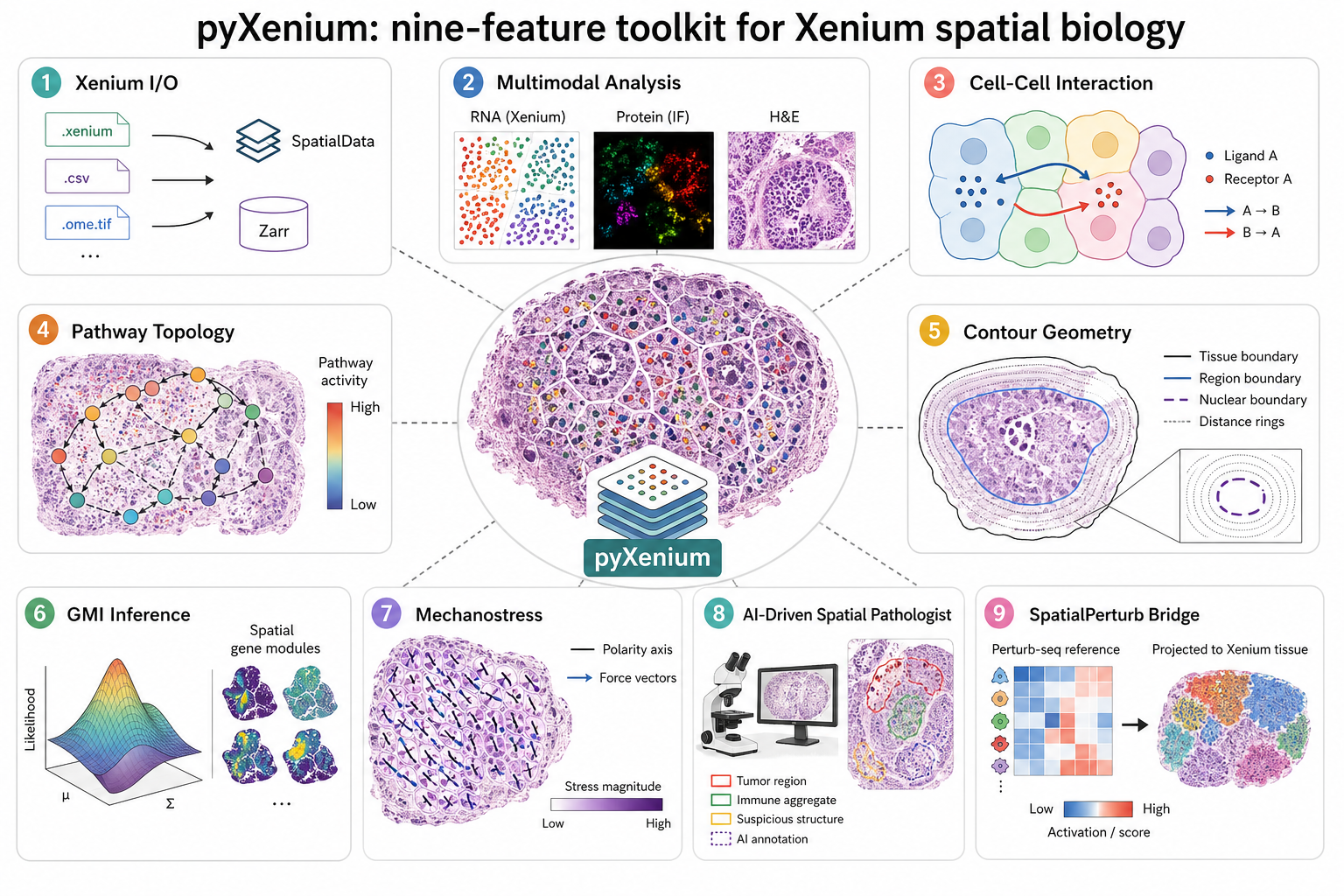
nine feature areas for Xenium I/O, multimodal analysis, CCI, pathway topology, contour geometry, GMI inference, mechanostress analysis, and external workflow bridges.
PyPI · Read the Docs · GitHub · Changelog · Releases
Figure 1. pyXenium summarizes Xenium I/O, multimodal analysis, spatial topology, contour-aware inference, mechanostress analysis, and external workflow bridges in one submission-ready overview.
pyXenium is a Python toolkit for 10x Genomics Xenium organized around nine major sections.
| Section | Canonical entry | Start here |
|---|---|---|
| Xenium I/O | pyXenium.io |
Xenium data loading guide |
| Multimodal Analysis | pyXenium.multimodal |
Multimodal overview |
| Cell-Cell Interaction | pyXenium.cci |
CCI tutorial hub |
| Pathway Topology | pyXenium.pathway |
Pathway tutorial |
| Contour Geometry | pyXenium.contour |
Contour tutorial hub |
| GMI Inference | pyXenium.gmi |
Contour GMI guide |
| Mechanostress | pyXenium.mechanostress |
Mechanostress tutorial |
| AI-Driven Spatial Pathologist | pyXenium.spatho |
RTD bridge guide |
| SpatialPerturb Bridge | pyXenium.perturb |
SpatialPerturb bridge guide |
Legacy compatibility entry points under pyXenium.analysis, pyXenium.validation, and
pyXenium.io.load_xenium_gene_protein(...) remain importable, but new code should target the
canonical pyXenium namespaces above. The spatho and SpatialPerturb workflows are installed
and run separately; pyXenium does not vendor them or add them as core runtime dependencies.
- Current repository version:
0.4.5 - Package index: PyPI
- Documentation site: pyxenium.readthedocs.io
- Canonical build status: GitHub Actions CI
- Supported Python:
>=3.8 - License: GNU AGPL v3.0 only
- Maintained by:
SPATHO AB
pip install pyXenium==0.4.5For local development:
git clone https://github.com/hutaobo/pyXenium
cd pyXenium
pip install -e ".[dev]"For documentation work:
pip install -e ".[docs]"For the optional SpatialPerturb Bridge runtime on Python 3.9+:
pip install -e ".[perturb]"from pyXenium.io import read_xenium
slide = read_xenium("/path/to/xenium_export", as_="slide", prefer="zarr")XeniumSlide is the only canonical in-memory object in pyXenium. Legacy
alias-based I/O entrypoints have been removed.
To build Atera-style learning stores with contour-segmented H&E crops:
pyxenium slide build-atera --atera-root Y:\long\10X_datasets\Xenium\Atera --output-root D:\GitHub\stGPT\outputs\xenium_slides\ateraThis writes one xenium_slide.zarr per case plus slide_manifest.json, qc_report.json, cell_to_contour.parquet, structure_assignments.csv, and contour_patches_manifest.json. Raw Xenium outs directories are read only; per-cell H&E patches are not generated.
from pyXenium.multimodal import load_rna_protein_anndata
adata = load_rna_protein_anndata(
base_path="/path/to/xenium_export",
prefer="auto",
)from pyXenium.contour import expand_contours
expand_contours(
slide,
contour_key="protein_cluster_contours",
distance=25.0,
mode="voronoi",
)from pyXenium.gmi import ContourGmiConfig, run_atera_breast_contour_gmi
result = run_atera_breast_contour_gmi(
dataset_root="/path/to/WTA_Preview_FFPE_Breast_Cancer_outs",
output_dir="pyxenium_gmi_outputs/atera_s1_s5",
config=ContourGmiConfig(feature_count=500, spatial_feature_count=100),
)pyXenium.gmi is a canonical beta surface: the API is public, while biological interpretation
should be checked with the bundled controls, cross-validation, and PDC Slurm reproducibility workflow.
The Atera S1/S5 validation completed on PDC Dardel in the v0.4.1 release, supporting an
S5/DCIS RNA program led by NIBAN1 and SORL1 under the QC20 primary model.
from pyXenium.io import read_xenium
from pyXenium.mechanostress import MechanostressConfig, run_mechanostress_workflow
slide = read_xenium("/path/to/xenium_export", as_="slide", include_boundaries=True)
result = run_mechanostress_workflow(
slide,
output_dir="pyxenium_mechanostress_outputs/sample_1",
config=MechanostressConfig(),
)For cohorts organized as one Xenium sample per directory, use:
from pyXenium.mechanostress import MechanostressConfig, run_mechanostress_cohort
cohort = run_mechanostress_cohort(
"/path/to/headandneckSCC",
output_dir="pyxenium_mechanostress_outputs/hnscc",
config=MechanostressConfig(coupling_genes=("KRT5", "EPCAM")),
)pyXenium.mechanostress is a canonical beta surface for extracting mechanical stress
signals from Xenium morphology and spatial cell context.
AI-Driven Spatial Pathologist
is an external workflow layer for AI-assisted pathology review around Xenium-scale spatial
transcriptomics. Its public Python package and CLI are named spatho.
pip install -U spatho
spatho init-workflow --organ breast --case-name breast_case_01 --dataset-root /path/to/Xenium_outs --output workflow.json
spatho doctor --config workflow.json
spatho run --config workflow.jsonIn pyXenium, this is documented as an optional external workflow bridge rather than a new
pyXenium.spatho namespace.
The handoff is possible because XeniumSlide keeps the cell table, transcript points,
cell/nucleus boundaries, H&E image metadata, and slide-native organization together
for downstream tools, with an optional XeniumSlide.to_spatialdata() bridge for users
who separately install that ecosystem.
XeniumSlide is inspired by the container ideas popularized by
SpatialData, while pyXenium now ships a
fully rewritten independent slide model and Zarr store implementation with no core runtime
dependency on the spatialdata package. If you build on the bridge or the design lineage,
please also cite Marconato et al., 2024.
SpatialPerturb is an external workflow package
for combining spatial transcriptomics with Perturb-seq references. pyXenium exposes a lightweight
pyXenium.perturb bridge that writes a handoff JSON and stable external CLI commands without
vendoring the SpatialPerturb algorithms.
from pyXenium.perturb import SpatialPerturbBridgeConfig, write_spatialperturb_handoff
spec = write_spatialperturb_handoff(
SpatialPerturbBridgeConfig(
xenium_path="/path/to/Xenium_outs",
output_dir="spatialperturb_reports/breast_case_01",
cell_group_path="/path/to/cell_groups.csv",
roi_geojson_path="/path/to/xenium_explorer_annotations.geojson",
sample_name="breast_case_01",
),
"spatialperturb_bridge.json",
)
print(spec["command_text"]["run_reference_benchmark"])SpatialPerturb Bridge scores mean Perturb-seq-derived program similarity projected onto Xenium tissue. They do not mean the tissue cell contains the corresponding knockout, guide, or drug perturbation.
The docs now separate the nine feature sections from the main reading paths:
Start here: pyxenium.readthedocs.io
This project is licensed under the GNU Affero General Public License v3.0 only.
It is maintained by SPATHO AB.
Copyright remains with Taobo Hu and other contributors where applicable.